Unlocking the Biocatalytic Efficiency of Factor-Independent Urate Hydroxylase from Bacillus subtilis in Petrochemical Hydrocarbon Detoxification

- Toxicity Analysis,
- Petrochemical Hydrocarbons,
- Computational Biodegradation,
- Factor Independent Urate Hydroxylase,
- Biocatalytic Pathway Analysis
- PTM ...More
Copyright (c) 2026 SCHQ

This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License.
Abstract
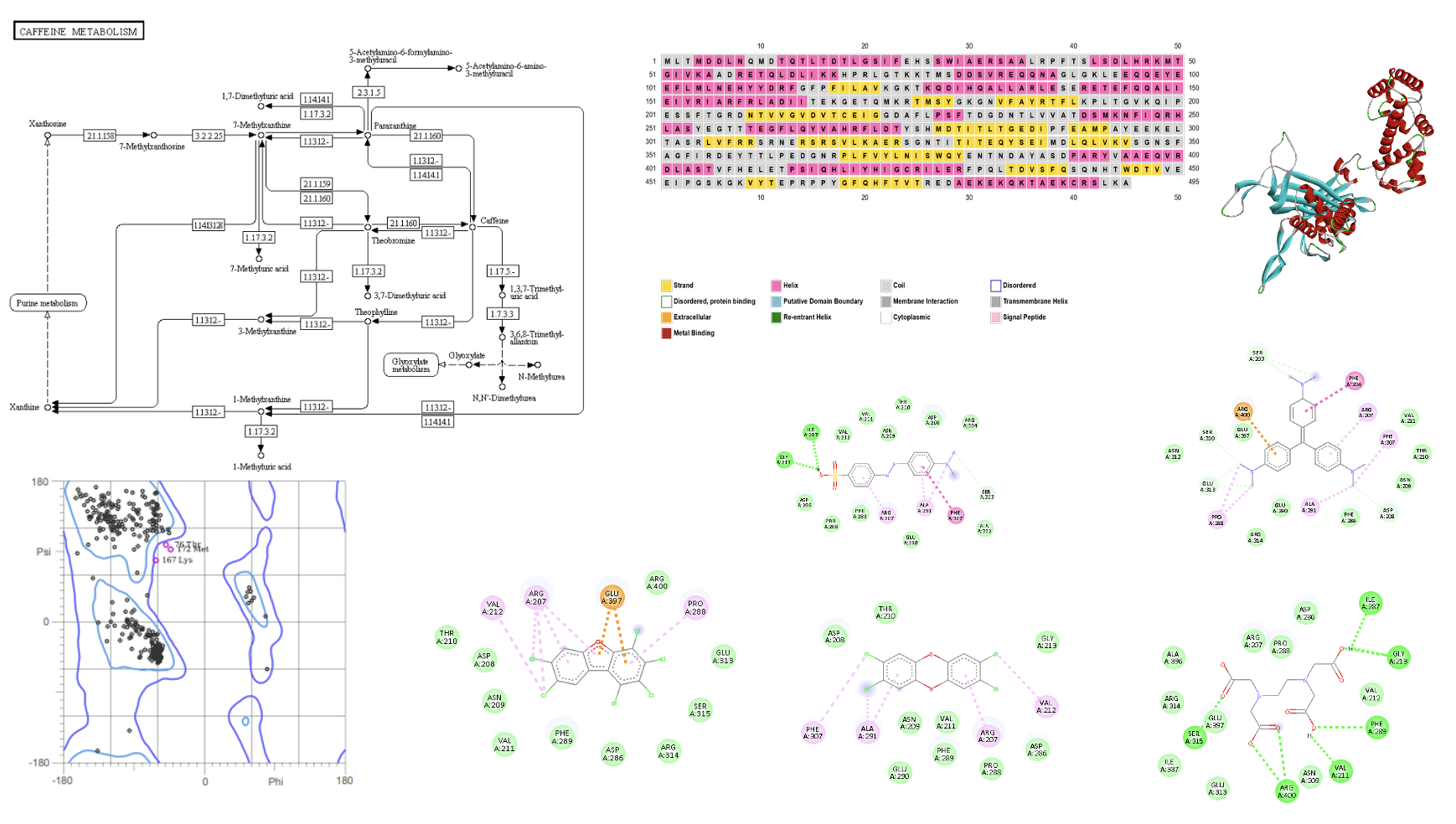
Petrochemical hydrocarbons are toxic, persistent environmental pollutants that pose serious ecological and health risks. Bioremediation using microbial enzymes offers a sustainable and effective alternative for their degradation. This study presents an in-silico analysis of Factor-independent urate hydroxylase from Bacillus subtilis to evaluate its potential in hydrocarbon biodegradation. The enzyme’s physicochemical properties revealed moderate stability and hydrophilicity, favoring activity in aqueous environments. Post-translational modification analysis predicted multiple regulatory sites, suggesting adaptability to environmental conditions. Structural modeling and validation confirmed a high-quality 3D structure suitable for molecular docking. Nine petrochemical hydrocarbons were selected for virtual screening. Docking results showed strong binding affinities, particularly with 1,2,3,4,7,8-hexachlorodibenzofuran (-7.4 kcal/mol), crystal violet (-7.3 kcal/mol), and dioxins, with key residues (e.g., ARG207, VAL211, PHE289) mediating interactions. Toxicity predictions indicated high neurotoxicity and hepatotoxicity among the compounds, highlighting the urgency for effective remediation tools. The study concludes that Factor-independent urate hydroxylase demonstrates promising interaction with harmful hydrocarbons and play a key role in microbial bioremediation. These computational findings provide a foundation for future experimental validation and potential application in cleaning up petrochemical-contaminated environments.
References
- BAI, Q., MA, J., LIU, S., XU, T., BANEGAS-LUNA, A. J., PÉREZ-SÁNCHEZ, H., TIAN, Y., HUANG, J., LIU, H., YAO, X. J. C. & JOURNAL, S. B. 2021. WADDAICA: A webserver for aiding protein drug design by artificial intelligence and classical algorithm. 19, 3573-3579.
- BANERJEE, P., ECKERT, A. O., SCHREY, A. K. & PREISSNER, R. 2018. ProTox-II: a webserver for the prediction of toxicity of chemicals. Nucleic acids research, 46, W257-W263.
- BUCHAN, D. W. & JONES, D. T. 2019. The PSIPRED protein analysis workbench: 20 years on. Nucleic acids research, 47, W402-W407.
- CHANG, K.-F., FANG, G.-C., CHEN, J.-C. & WU, Y.-S. 2006. Atmospheric polycyclic aromatic hydrocarbons (PAHs) in Asia: a review from 1999 to 2004. Environmental pollution, 142, 388-396.
- COUTO, S. R. & HERRERA, J. L. T. 2006. Industrial and biotechnological applications of laccases: a review. Biotechnology advances, 24, 500-513.
- DALLAKYAN, S. & OLSON, A. J. 2014. Small-molecule library screening by docking with PyRx. Chemical biology: methods and protocols. Springer.
- HARITASH, A. & KAUSHIK, C. 2009. Biodegradation aspects of polycyclic aromatic hydrocarbons (PAHs): a review. Journal of hazardous materials, 169, 1-15.
- HYNDMAN, D., LIU, S. & MINER, J. N. 2016. Urate handling in the human body. Current rheumatology reports, 18, 34.
- KEENAN, R. T. The biology of urate. Seminars in arthritis and rheumatism, 2020. Elsevier, S2-S10.
- KIM, K.-H., JAHAN, S. A., KABIR, E. & BROWN, R. J. 2013. A review of airborne polycyclic aromatic hydrocarbons (PAHs) and their human health effects. Environment international, 60, 71-80.
- KIM, S., THIESSEN, P. A., BOLTON, E. E., CHEN, J., FU, G., GINDULYTE, A., HAN, L., HE, J., HE, S. & SHOEMAKER, B. A. 2016. PubChem substance and compound databases. Nucleic acids research, 44, D1202-D1213.
- KUBIČKOVÁ, I. & KUBIČKA, D. 2010. Utilization of triglycerides and related feedstocks for production of clean hydrocarbon fuels and petrochemicals: a review. Waste and Biomass Valorization, 1, 293-308.
- MACEK, B., MANN, M. & OLSEN, J. V. 2009. Global and site-specific quantitative phosphoproteomics: principles and applications. Annual review of pharmacology and toxicology, 49, 199-221.
- MATAR, S. & HATCH, L. F. 2001. Chemistry of petrochemical processes, Elsevier.
- NAVEED, M., NAVEED, R., AZIZ, T., AZEEM, A., AFZAL, M., WASEEM, M., ALHARBI, M., ALSHAMMARI, A., ALASMARI, A. F. & ALBEKAIRI, T. H. J. B. 2024a. Biodegradation of PVCs through in-vitro identification of Bacillus albus and computational pathway analysis of ABH enzyme. 35, 451-468.
- NAVEED, M., NAVEED, R., AZIZ, T., IQBAL, F., HASSAN, A., SALEEM, A., WASEEM, M., RAHMAN, S. U., ALHARBI, M., ALSHAMMARI, A. J. B. C. & BIOREFINERY 2025. Exploring the potential application of peroxidase enzyme from Acinetobacter baumannii as an eco-friendly agent for the bioremediation of the highly noxious pyrethroid compounds through molecular docking analysis. 15, 155-173.
- NAVEED, M., SALAH UD DIN, M., AZIZ, T., JAVED, T., MIRAJ KHAN, S., NAVEED, R., ALI KHAN, A. & ALHARBI, M. J. Z. F. N. C. 2024b. Comparative analysis among the degradation potential of enzymes obtained from Escherichia coli against the toxicity of sulfur dyes through molecular docking. 79, 221-234.
- NAVEED, M., SALEEM, A., AZIZ, T., NAVEED, R., DIN, M. S. U., SHABBIR, M. A., KHAN, A. A., ALAMRI, A. S., ALSANIE, W. F., ALHOMRANI, M. J. J. O. C. B. & CHEMISTRY 2024c. Differentially Expressed Genes in Lung Cancer: Molecular Docking of Hypoxanthine Guanine-Phosphoribosyl Transferase with Dibutyl Phthalate. 23, 1375-1387.
- NAVEED, M., SHABBIR, M. A., AZIZ, T., SALEEM, A., NAVEED, R., KHAN, A. A., HAQ, T. U., ALHARBI, M., ALSHAMMARI, A. & ALASMARI, A. F. J. A. B. P. 2023. In silico explorations of bacterial mercuric reductase as an ecofriendly bioremediator for noxious mercuric intoxications. 70, 661-669.
- PÉREZ‐PANTOJA, D., DONOSO, R., AGULLÓ, L., CÓRDOVA, M., SEEGER, M., PIEPER, D. H. & GONZÁLEZ, B. 2012. Genomic analysis of the potential for aromatic compounds biodegradation in Burkholderiales. Environmental microbiology, 14, 1091-1117.
- ROH, C., CHOI, K.-Y., PANDEY, B. P. & KIM, B.-G. 2009. Hydroxylation of daidzein by CYP107H1 from Bacillus subtilis 168. Journal of Molecular Catalysis B: Enzymatic, 59, 248-253.
- SCHOCH, C. L., CIUFO, S., DOMRACHEV, M., HOTTON, C. L., KANNAN, S., KHOVANSKAYA, R., LEIPE, D., MCVEIGH, R., O’NEILL, K. & ROBBERTSE, B. 2020. NCBI Taxonomy: a comprehensive update on curation, resources and tools. Database, 2020, baaa062.
- SCHWEDE, T., KOPP, J., GUEX, N. & PEITSCH, M. C. 2003. SWISS-MODEL: an automated protein homology-modeling server. Nucleic acids research, 31, 3381-3385.
- SHI, Y., ZHANG, X., YANG, Y., CAI, T., PENG, C., WU, L., ZHOU, L., HAN, J., MA, M. & ZHU, W. 2023. D3CARP: a comprehensive platform with multiple-conformation based docking, ligand similarity search and deep learning approaches for target prediction and virtual screening. Computers in Biology and Medicine, 164, 107283.
- SINGH, P., JAIN, R., SRIVASTAVA, N., BORTHAKUR, A., PAL, D., SINGH, R., MADHAV, S., SRIVASTAVA, P., TIWARY, D. & MISHRA, P. K. 2017. Current and emerging trends in bioremediation of petrochemical waste: a review. Critical Reviews in Environmental Science and Technology, 47, 155-201.
- VIRK, G. K. 2022. Blood based biomarkers for the identification of Alzheimer’s disease using proteomics approaches. University of New South Wales (Australia).
- WANG, D., ZENG, S., XU, C., QIU, W., LIANG, Y., JOSHI, T. & XU, D. 2017. MusiteDeep: a deep-learning framework for general and kinase-specific phosphorylation site prediction. Bioinformatics, 33, 3909-3916.
- WATTAM, A. R., ABRAHAM, D., DALAY, O., DISZ, T. L., DRISCOLL, T., GABBARD, J. L., GILLESPIE, J. J., GOUGH, R., HIX, D. & KENYON, R. 2014. PATRIC, the bacterial bioinformatics database and analysis resource. Nucleic acids research, 42, D581-D591.
- WILCKE, W. 2000. Synopsis polycyclic aromatic hydrocarbons (PAHs) in soil—a review. Journal of plant nutrition and soil science, 163, 229-248.